|
8/13/2023 0 Comments Calculating formal charge in atoms 
The utility of OFraMP is illustrated using the anti-cancer agent paclitaxel and a dendrimer used in organic semiconductor devices. OFraMP also allows users to manually alter interaction parameters and automates the submission of missing substructures to the ATB in order to generate parameters for atoms in environments not represented in the existing database. The user then selects the most appropriate match. Adjacent matching atoms are combined into progressively larger matched sub-structures. Atoms are considered within the context of an extended local environment (buffer region) with the degree of similarity between an atom in the target molecule and that in the proposed match controlled by varying the size of the buffer region. OFraMP identifies and compares alternative molecular fragments from the ATB database, which contains over 890,000 pre-parameterized molecules, using a novel hierarchical matching procedure. OFraMP is a web application for assigning atomic interaction parameters to large molecules by matching sub-fragments within the target molecule to equivalent sub-fragments within the Automated Topology Builder (ATB, atb.uq.edu.au) database. Journal of Computer-Aided Molecular DesignĪn Online tool for Fragment-based Molecule Parametrization (OFraMP) is described. 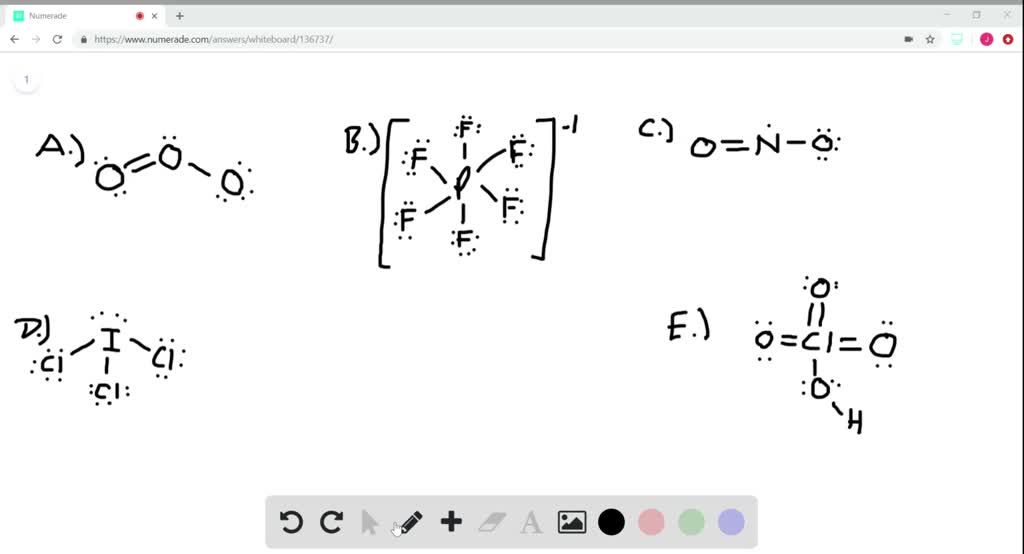
OFraMP: a fragment-based tool to facilitate the parametrization of large molecules 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
